Lesson 5: Why Hierarchical Modelling Beats Maximum Likelihood¶
The problem with individual fitting¶
Maximum-likelihood estimation (MLE) and MAP estimation fit each participant independently. With hundreds of trials per subject this works fine, but in practice most psychophysics experiments give you 100–250 trials per condition. At those trial counts individual MLE/MAP estimates are:
Noisy — refit the same subject on a different random half of their data and you get a substantially different answer.
Biased at the boundaries — with few trials, the likelihood surface is broad and the optimiser can land at extreme values. Noise parameters (\(\nu\)) may converge to near-zero (the model “explains” every trial perfectly by overfitting) or explode to very large values (flat psychometric curve, no discrimination at all). Lapse-rate parameters can rail at 0 or 1. These boundary estimates are not meaningful — they reflect the instability of the optimisation, not the participant’s true noise level.
Worse for complex models — every additional parameter increases the volume of the parameter space that the optimiser must search. The most theoretically interesting models (KLW with prior \(\sigma_0\), FlexibleNoise with spline coefficients) are exactly the ones that suffer most, because they have more parameters per subject and more opportunities for the likelihood to be flat.
Hierarchical Bayesian estimation avoids all three problems. By sharing statistical strength across participants, the group prior acts as an adaptive regulariser: subjects with extreme or unreliable data are pulled toward the population, while well-identified subjects are left alone.
Empirical demonstration: split-half reliability¶
The gold standard for measurement quality is split-half reliability: split each participant’s trials into two random halves, fit each half separately, and correlate the parameter estimates across halves. A reliable method produces the same answer from both halves; an unreliable one does not.
We compare:
Method |
What it does |
Regularisation |
|---|---|---|
MLE |
|
None — pure maximum likelihood |
Hierarchical Bayes |
|
Adaptive group prior, posterior averaging |
at increasing trial counts (25, 50, 108 per half). The Garcia et al. magnitude data have 216 trials per subject, so 108 per half is the natural split.
[1]:
import warnings; warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from bauer.utils.data import load_garcia2022
from bauer.models import MagnitudeComparisonModel
df_mag = load_garcia2022(task='magnitude')
subjects = df_mag.index.get_level_values('subject').unique()
print(f"Subjects: {len(subjects)}, trials per subject: "
f"{df_mag.groupby(level='subject').size().iloc[0]}")
# ── Split each subject's trials into two random halves ───────────────────────
rng = np.random.default_rng(42)
half_a_rows, half_b_rows = [], []
for subj in subjects:
mask = df_mag.index.get_level_values('subject') == subj
subj_iloc = np.where(mask)[0]
perm = rng.permutation(len(subj_iloc))
mid = len(subj_iloc) // 2
half_a_rows.extend(subj_iloc[perm[:mid]])
half_b_rows.extend(subj_iloc[perm[mid:]])
half_a_full = df_mag.iloc[half_a_rows]
half_b_full = df_mag.iloc[half_b_rows]
print(f"Half A: {len(half_a_full)} trials, Half B: {len(half_b_full)} trials")
Subjects: 64, trials per subject: 216
Half A: 6912 trials, Half B: 6912 trials
Split-half analysis: 10 random splits \(\times\) 5 trial counts¶
For each trial count (25, 50, 108) we:
Randomly split each subject’s data into two halves
Subsample k trials per half
Fit MLE and hierarchical Bayes on each half
Correlate the parameter estimates across halves
We repeat this 10 times with different random splits to get a stable estimate of reliability. Three correlation metrics are shown: Spearman \(\rho\), Pearson \(r\), and \(R^2\).
[2]:
from scipy.stats import pearsonr
def subsample(df, k):
return df.groupby(level='subject').head(k)
def split_data(df_mag, seed):
# Random split of each subject's trials into two halves
rng = np.random.default_rng(seed)
a_rows, b_rows = [], []
for subj in df_mag.index.get_level_values('subject').unique():
mask = df_mag.index.get_level_values('subject') == subj
iloc = np.where(mask)[0]
perm = rng.permutation(len(iloc))
mid = len(iloc) // 2
a_rows.extend(iloc[perm[:mid]])
b_rows.extend(iloc[perm[mid:]])
return df_mag.iloc[a_rows], df_mag.iloc[b_rows]
def fit_mle(data):
model = MagnitudeComparisonModel(paradigm=data, fit_seperate_evidence_sd=True)
return model.fit_map_individual(data=data, flat_prior=True)
def fit_hierarchical(data, draws=500, tune=500, chains=2):
model = MagnitudeComparisonModel(paradigm=data, fit_seperate_evidence_sd=True)
model.build_estimation_model(data=data, hierarchical=True)
idata = model.sample(draws=draws, tune=tune, chains=chains, progressbar=False)
n_subj = len(data.index.unique(level='subject'))
n1 = idata.posterior['n1_evidence_sd'].values.reshape(-1, n_subj).mean(0)
n2 = idata.posterior['n2_evidence_sd'].values.reshape(-1, n_subj).mean(0)
return pd.DataFrame({'n1_evidence_sd': n1, 'n2_evidence_sd': n2},
index=pd.Index(data.index.unique(level='subject'), name='subject'))
trial_counts = [25, 50, 108]
n_splits = 3
methods = {'MLE': fit_mle, 'Hierarchical Bayes': fit_hierarchical}
results = []
for split_i in range(n_splits):
half_a, half_b = split_data(df_mag, seed=split_i)
for k in trial_counts:
a_sub = subsample(half_a, k)
b_sub = subsample(half_b, k)
for method_name, fit_fn in methods.items():
est_a = fit_fn(a_sub)
est_b = fit_fn(b_sub)
# Also compute sum of both noise params
est_a['total_sd'] = est_a['n1_evidence_sd'] + est_a['n2_evidence_sd']
est_b['total_sd'] = est_b['n1_evidence_sd'] + est_b['n2_evidence_sd']
for param in ['n1_evidence_sd', 'n2_evidence_sd', 'total_sd']:
rho_p, _ = pearsonr(est_a[param], est_b[param])
# Count boundary-collapsed estimates (noise near zero)
n_zero_a = (est_a[param] < 0.01).sum()
n_zero_b = (est_b[param] < 0.01).sum()
results.append({
'split': split_i, 'k': k, 'method': method_name,
'parameter': param,
'Pearson r': rho_p, 'R\u00b2': rho_p**2,
'n_collapsed': n_zero_a + n_zero_b,
})
print(f"Split {split_i+1}/{n_splits} done")
results_df = pd.DataFrame(results)
print(f"\nTotal fits: {len(results_df)}")
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 5 seconds.
There was 1 divergence after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 5 seconds.
There were 6 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 9 seconds.
There were 16 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 13 seconds.
There were 9 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 28 seconds.
There were 5 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 22 seconds.
There were 2 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
Split 1/3 done
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 4 seconds.
There was 1 divergence after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 5 seconds.
There were 4 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 28 seconds.
There was 1 divergence after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 10 seconds.
There were 34 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 30 seconds.
There were 4 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 22 seconds.
There were 5 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Split 2/3 done
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 5 seconds.
There were 4 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 5 seconds.
There were 8 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 10 seconds.
There were 9 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 10 seconds.
There were 41 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 21 seconds.
There were 87 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 21 seconds.
There were 6 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Split 3/3 done
Total fits: 54
[3]:
# ── Reliability vs trial count: Pearson r and R² x 3 parameters ──────────────
metrics = ['Pearson r', 'R\u00b2']
params = ['n1_evidence_sd', 'n2_evidence_sd', 'total_sd']
param_labels = {'n1_evidence_sd': '\u03bd\u2081 (first option)',
'n2_evidence_sd': '\u03bd\u2082 (second option)',
'total_sd': '\u03bd\u2081 + \u03bd\u2082 (total)'}
pal = {'MLE': '#d73027', 'Hierarchical Bayes': '#4393c3'}
fig, axes = plt.subplots(len(metrics), len(params),
figsize=(5 * len(params), 3.5 * len(metrics)),
sharex=True, sharey='row')
for row, metric in enumerate(metrics):
for col, param in enumerate(params):
ax = axes[row, col]
sub = results_df[results_df['parameter'] == param]
for i_m, method in enumerate(['MLE', 'Hierarchical Bayes']):
d = sub[sub['method'] == method]
mean_by_k = d.groupby('k')[metric].mean()
se_by_k = d.groupby('k')[metric].std() / np.sqrt(n_splits)
offset = (i_m - 0.5) * 1.5 # slight x-offset to avoid overlap
ax.errorbar(mean_by_k.index + offset, mean_by_k,
yerr=1.96 * se_by_k,
fmt='o-', lw=2, ms=6, capsize=4, capthick=1.5,
color=pal[method], ecolor=pal[method], label=method)
ax.axhline(0, ls=':', c='gray', lw=1)
if row == 0:
ax.set_title(param_labels[param])
if row == len(metrics) - 1:
ax.set_xlabel('Trials per half')
if col == 0:
ax.set_ylabel(metric)
ax.legend(fontsize=8)
sns.despine(ax=ax)
plt.suptitle(f'Split-half reliability (mean \u00b1 95% CI over {n_splits} splits)',
fontsize=13, y=1.01)
plt.tight_layout()
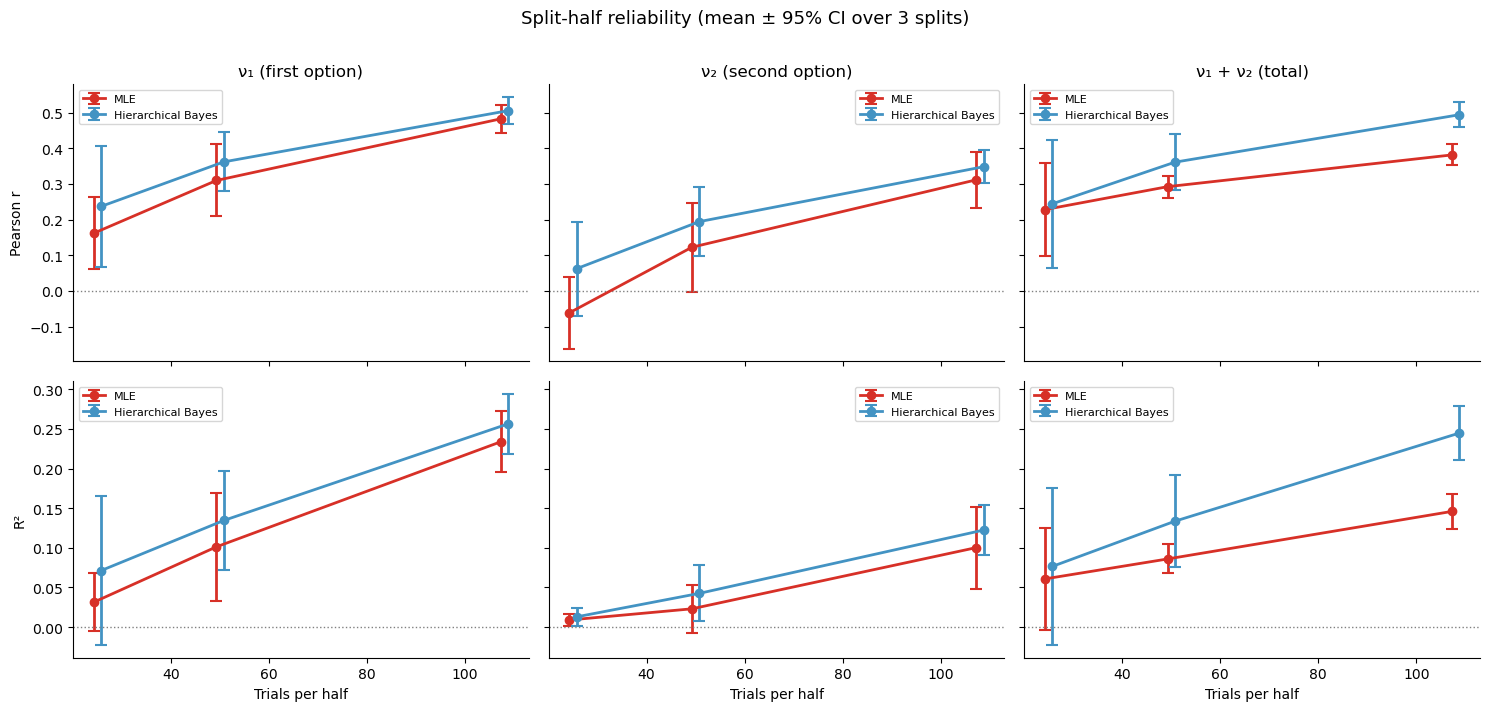
The boundary-collapse problem¶
Why does MLE do so poorly on Pearson \(r\) (which measures linear agreement) even when Spearman \(\rho\) (which only checks rank order) looks acceptable? The answer is boundary collapse: with few trials, the MLE optimiser can push noise parameters \(\nu_1\) or \(\nu_2\) to near-zero, meaning the model “explains” every trial perfectly by overfitting the noise away. These zero-estimates are meaningless — they do not reflect a participant with perfect perception, they reflect an optimiser that ran out of data.
The plot below shows how many subjects (out of \(N \times 2\) halves) have \(\nu < 0.01\) at each trial count:
[4]:
# ── Boundary collapse diagnostic ─────────────────────────────────────────────
fig, axes = plt.subplots(1, 3, figsize=(14, 4), sharey=True)
for col, param in enumerate(params):
ax = axes[col]
sub = results_df[results_df['parameter'] == param]
for i_m, method in enumerate(['MLE', 'Hierarchical Bayes']):
d = sub[sub['method'] == method]
mean_col = d.groupby('k')['n_collapsed'].mean()
se_col = d.groupby('k')['n_collapsed'].std() / np.sqrt(n_splits)
offset = (i_m - 0.5) * 1.5
ax.errorbar(mean_col.index + offset, mean_col,
yerr=1.96 * se_col,
fmt='o-', lw=2, ms=6, capsize=4, capthick=1.5,
color=pal[method], ecolor=pal[method], label=method)
ax.axhline(0, ls=':', c='gray', lw=1)
ax.set_title(param_labels[param])
ax.set_xlabel('Trials per half')
if col == 0:
ax.set_ylabel('Subjects with \u03bd < 0.01\n(out of 2 \u00d7 N halves)')
ax.legend(fontsize=8)
sns.despine(ax=ax)
plt.suptitle('Boundary collapse: MLE pushes noise to zero when data are scarce',
fontsize=12, y=1.02)
plt.tight_layout()
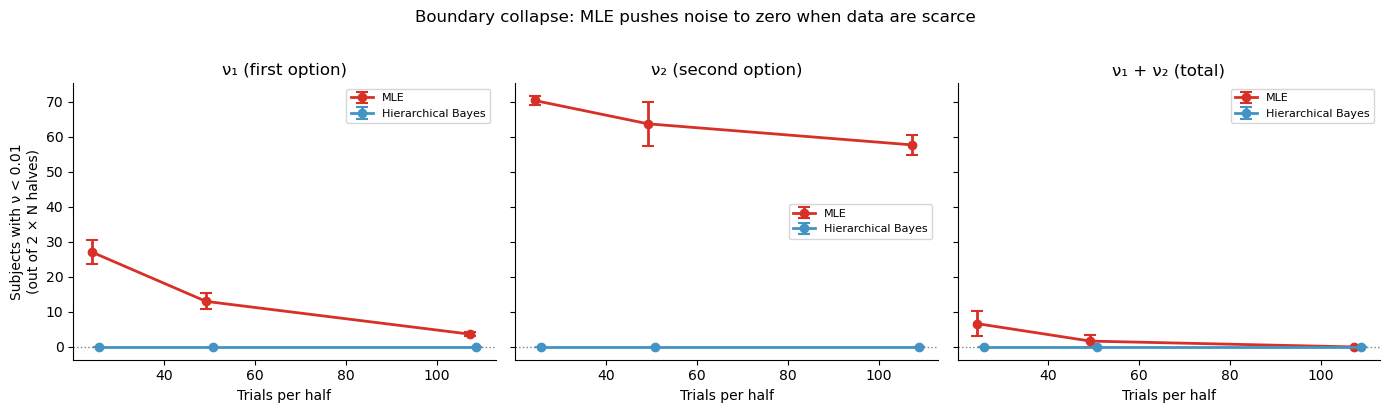
What this means in practice¶
The plot shows what every psychophysics researcher should know but few textbooks make explicit:
Individual MAP/MLE is unreliable at typical trial counts. At 50–100 trials per subject the split-half correlation can be near zero — the estimates are dominated by sampling noise and tell you almost nothing about the participant.
Hierarchical Bayes is dramatically more reliable. The group prior acts as adaptive regularisation: extreme/noisy estimates are pulled toward the group mean, which is exactly what reduces split-half variability. The posterior mean is a shrinkage estimator — the optimal bias-variance trade-off.
The advantage is largest when you need it most — at low trial counts and for complex models with many parameters. The KLW model (lesson 2) and the FlexibleNoise model (lesson 4) have 4+ parameters per subject; individual MLE would be hopeless without hundreds of trials, but hierarchical fitting works with the trial counts we actually have.
Why MAP/MLE fails: the bias-variance trade-off¶
MAP gives the single point with highest posterior density — for a flat prior this is just maximum likelihood. At low trial counts:
The likelihood surface is flat in some directions, so the optimiser lands at an arbitrary point in a large region of near-equal likelihood.
Boundary effects distort estimates: noise parameters can converge to near-zero (overfitting a lucky sample) or blow up.
There is no uncertainty quantification: you get a point estimate with no error bar, so you cannot distinguish a well-identified subject from a poorly-identified one.
Hierarchical Bayes solves all three: the group prior tilts the likelihood surface toward sensible values, the posterior is a full distribution (not a point), and the amount of shrinkage is automatically calibrated per subject.
Rule of thumb¶
If you have < 200 trials per subject per condition, always use hierarchical fitting. If you have < 100, individual fitting is essentially meaningless for models with more than one or two parameters. bauer makes hierarchical fitting the default for exactly this reason.
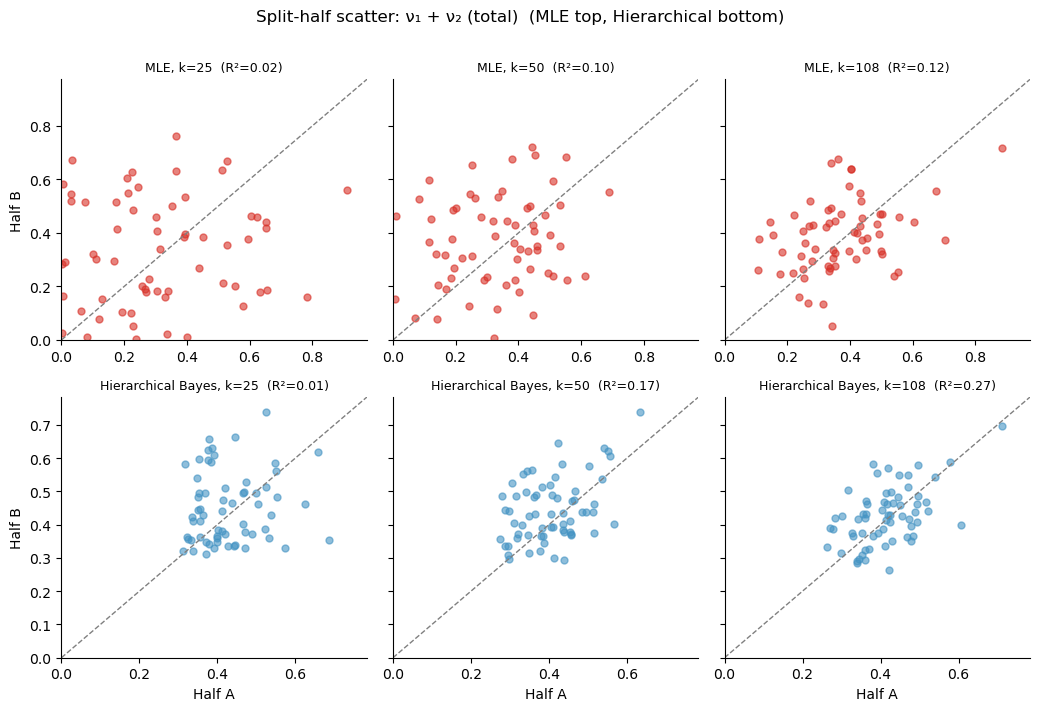
[5]:
# ── Scatter: half A vs half B at every trial count, all 3 params ─────────────
half_a, half_b = split_data(df_mag, seed=0)
scatter_colors = {'MLE': '#d73027', 'Hierarchical Bayes': '#4393c3'}
for param, param_label in param_labels.items():
n_k = len(trial_counts)
fig, axes = plt.subplots(2, n_k, figsize=(3.5 * n_k, 7), sharey='row', sharex='row')
for col, k in enumerate(trial_counts):
a_sub = subsample(half_a, k)
b_sub = subsample(half_b, k)
ests = {}
for name, fn in methods.items():
ea, eb = fn(a_sub), fn(b_sub)
ea['total_sd'] = ea['n1_evidence_sd'] + ea['n2_evidence_sd']
eb['total_sd'] = eb['n1_evidence_sd'] + eb['n2_evidence_sd']
ests[name] = (ea, eb)
for row, method in enumerate(['MLE', 'Hierarchical Bayes']):
ax = axes[row, col]
est_a, est_b = ests[method]
rho_p, _ = pearsonr(est_a[param], est_b[param])
ax.scatter(est_a[param], est_b[param], s=25, alpha=.6,
color=scatter_colors[method])
lims = [0, max(est_a[param].max(), est_b[param].max()) * 1.1]
ax.plot(lims, lims, '--', color='gray', lw=1)
ax.set_xlim(lims); ax.set_ylim(lims)
ax.set_title(f'{method}, k={k} (R\u00b2={rho_p**2:.2f})', fontsize=9)
if col == 0:
ax.set_ylabel(f'Half B')
if row == 1:
ax.set_xlabel(f'Half A')
sns.despine(ax=ax)
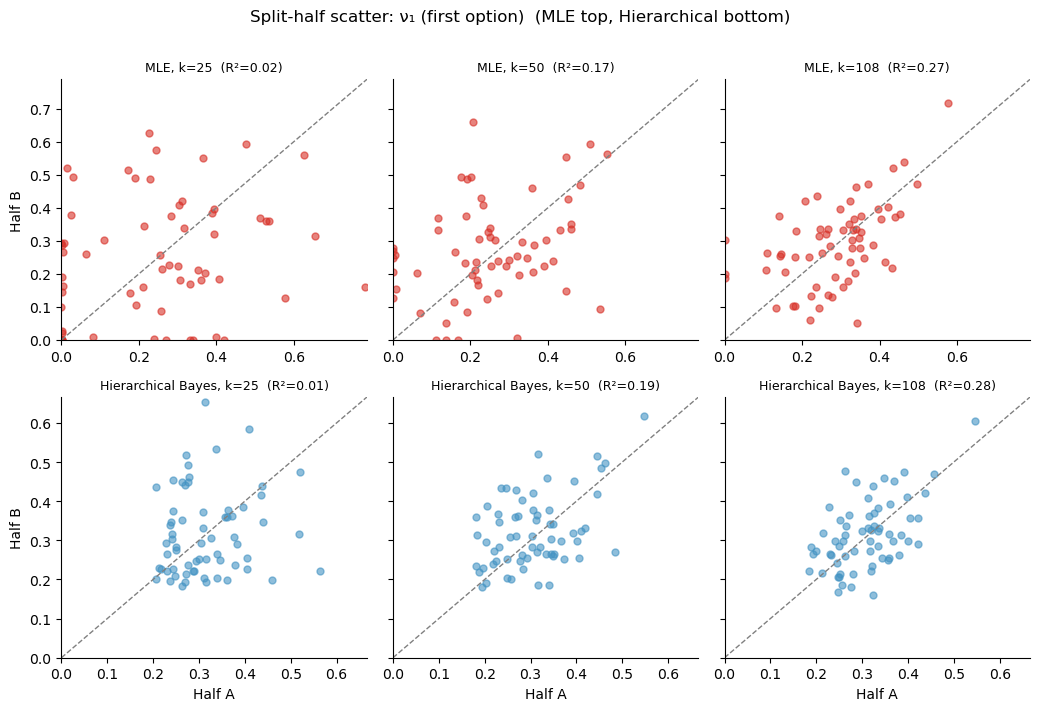
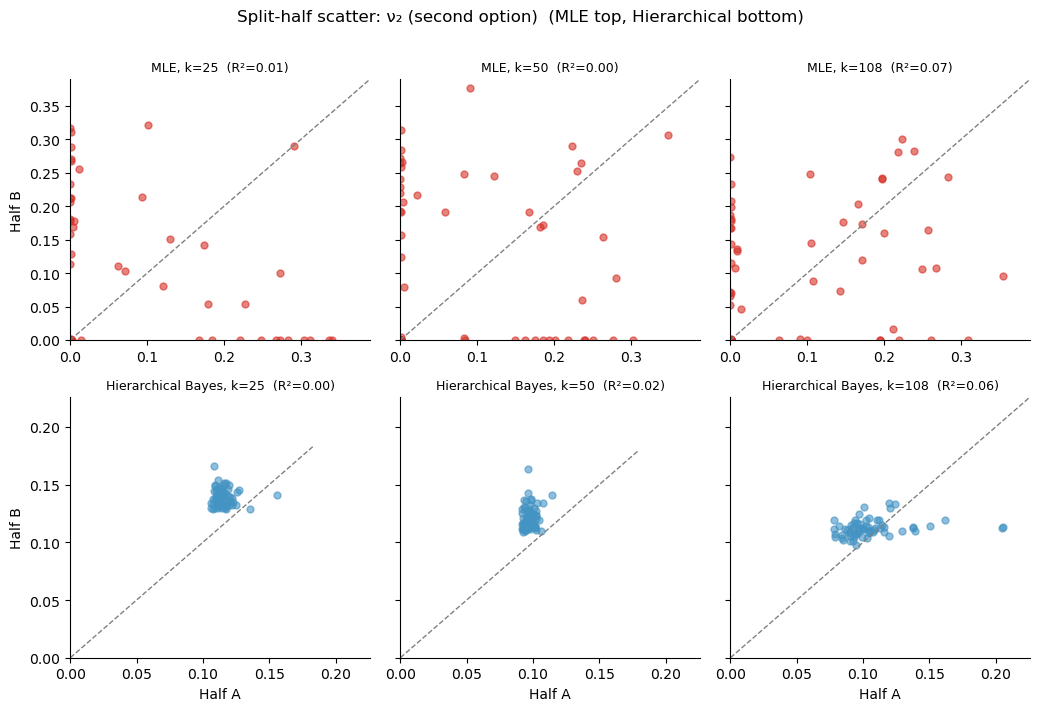
plt.suptitle(f'Split-half scatter: {param_label} (MLE top, Hierarchical bottom)',
fontsize=12, y=1.01)
plt.tight_layout()
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 5 seconds.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 5 seconds.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 12 seconds.
There were 31 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 10 seconds.
There were 13 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 29 seconds.
There were 8 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 22 seconds.
There were 309 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 5 seconds.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 5 seconds.
There was 1 divergence after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 9 seconds.
There were 5 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 10 seconds.
There were 8 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 29 seconds.
There were 45 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 27 seconds.
There were 53 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 5 seconds.
There was 1 divergence after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 5 seconds.
There was 1 divergence after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 10 seconds.
There were 24 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 10 seconds.
There were 10 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 33 seconds.
There were 6 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (2 chains in 2 jobs)
NUTS: [n1_evidence_sd_mu_untransformed, n1_evidence_sd_sd, n1_evidence_sd_offset, n2_evidence_sd_mu_untransformed, n2_evidence_sd_sd, n2_evidence_sd_offset]
Sampling 2 chains for 500 tune and 500 draw iterations (1_000 + 1_000 draws total) took 24 seconds.
There were 7 divergences after tuning. Increase `target_accept` or reparameterize.
We recommend running at least 4 chains for robust computation of convergence diagnostics
The rhat statistic is larger than 1.01 for some parameters. This indicates problems during sampling. See https://arxiv.org/abs/1903.08008 for details
The effective sample size per chain is smaller than 100 for some parameters. A higher number is needed for reliable rhat and ess computation. See https://arxiv.org/abs/1903.08008 for details
Take-aways¶
Never use individual MLE/MAP for cognitive models at typical psychophysics trial counts. The estimates are unreliable and any downstream correlation (e.g. with neural data or clinical scores) will be attenuated toward zero.
Hierarchical Bayes is not just “nicer” — it is a prerequisite for valid individual- difference analyses at the trial counts we actually work with.
bauer defaults to hierarchical fitting (
hierarchical=True) for exactly this reason. Individual MAP (model.fit_map()) is available for quick sanity checks but should not be used for final inference.These results generalise beyond magnitude comparison: the noisier your model and the fewer your trials, the bigger the hierarchical advantage.